Benchmark REPLICA : exemples de parsing
Principe du parsing des sorties Tripoli-4 standard
Le parsing est effectué en deux temps :
un premier parcours rapide du fichier répérant un certain nombre de balises (nombre de bacthes requis, temps d’initialisation, de simulation,
NORMAL COMPLETION, etc) module
module scan.py. Ce parcours permet également de repérer les résultats dans la sortie, débutant parRESULTS ARE GIVENet terminant le plus souvent par'simulation time:'.le réel parsing des résultats, piloté par le module
parse.pyqui appelle la grammaire (utilisantpyparsing) et transformant le résultat en objets python standards (listes, dictionnaires, tableaux numpy).
Exemple de parsing, jeu de données fast_neutron.t4
Présentation du jeu de données
Le jeu de données est issu du benchmark REPLICA du programme SINBAD, dont la géométrie Tripoli-4 est représentée ci-dessous. Il s’agit ici de la simulation des neutrons rapides.

2 types de réponses sont attendus :
réactions dans des détecteurs
flux surfacique
Dans le cas des neutrons rapides on dispose de 3 détecteurs : 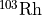 (
(RH103_IRDF85),  (
(IN115_IRDF85) et  (
(S32_IRDF85).
On fait ces mesures à différents emplacements. Le résultat final est constitué de :
10 réactions sur le
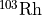 (10 frontières de volume différentes)
(10 frontières de volume différentes)3 réactions sur le

3 réactions sur le

2 flux surfaciques
Les emplacements des mesures sont représentés ci-dessous.

Étape 1 : scan puis parsing du résultat
On charge le module Parser. Le jeu de données est scanné automatiquement, tous les résultats de batches sont stockés dans la variable scan_res. On peut ensuite parser ceux qui nous intéressent.
Par défaut le résultat parsé sera le dernier batch, mais on pourrait en parser un autre.
Dernier batch  -1
-1
Scan du jeu de données
[1]:
from valjean.eponine.tripoli4.parse import Parser
[2]:
t4vv_replica_fast = 'fast_neutron.res'
t4p = Parser(t4vv_replica_fast)
INFO parse: Parsing fast_neutron.res
INFO scan: Edition batch (-1) different from current batch (50)
INFO scan: If no Edition batch keep current batch, else keep edition batch
INFO scan: Edition batch (-1) different from current batch (100)
INFO scan: If no Edition batch keep current batch, else keep edition batch
INFO parse: Successful scan in 0.016370 s
Parmi les membres de t4p il y a le résultat du scan. Certains paramètres accessibles à partir de ce résultat sont montrés ici :
parallèle ou mono-processeur (booléen)
les temps
[3]:
t4p.scan_res.normalend
[3]:
True
[4]:
t4p.scan_res.para
[4]:
False
[5]:
type(t4p.scan_res)
[5]:
valjean.eponine.tripoli4.scan.Scanner
[6]:
t4p.scan_res.times
[6]:
OrderedDict([('initialization_time', 12),
('simulation_time', {50: 4, 100: 9})])
Parsing du jeu de données
Le jeu de données peut être parser soit par numéro du batch quand on le connaît, grâce à la méthode parse_from_number, soit par son index dans la liste des batches scannés grâce à la méthode parse_from_index.
Le plus souvent c’est le dernier batch qui nous intéresse, le défaut de parse_from_index est donc -1.
Ces méthodes renvoient un ParseResult qui contient le batch parsé ainsi que les variables globales du jeu de données récupérées lors du scan (objet res).
Le ParseResult peut alors être transformé en Browser pour faciliter l’accès aux différents résultats grâce à la méthode to_browser.
[7]:
t4pres = t4p.parse_from_index()
INFO parse: Successful parsing in 0.175471 s
[8]:
len(t4pres.res), type(t4pres.res), list(t4pres.res.keys())
[8]:
(3, dict, ['list_responses', 'batch_data', 'run_data'])
[9]:
print('run_data:', t4pres.res['run_data'])
print('batch_data:', t4pres.res['batch_data'])
run_data: {'warnings': 1, 'errors': 0, 'number_of_tasks': 1, 'normal_end': True, 'required_batches': 100, 'partial': False, 't4_file': 'fast_neutron.res', 'initialization_time': 12}
batch_data: {'source_intensity': 81114.52, 'mean_weight_leak': {'score': 359.8768, 'sigma': 143.1067, 'sigma%': 39.76546}, 'simulation_time': 9, 'batch_number': 100, 'name': ''}
Le résultat stocké dans ParseResult est un dictionnaire.
'batch_data'correspond aux données spécifiques du batch (temps de simulation cumulé à la fin de celui-ci, numéro d’édition, intensité de la source, etc.)'run_data'correspond aux données globales du jeu de données (temps d’initialisation, les nombres d’erreurs et de warnings, le nom du jeu de données, etc.'list_responses'est la liste des résultats (liste de dictionnaires)
[10]:
print(type(t4pres.res['list_responses']))
print(type(t4pres.res['list_responses'][-1]))
<class 'list'>
<class 'dict'>
Les différents résultats sont stockés dans la liste dans l’ordre où ils apparaissent dans la sortie de TRIPOLI4, sous forme de dictionnaires.
Exemple court :
[11]:
print(t4pres.res['list_responses'][0])
{'response_function': 'REACTION', 'response_name': 'reaction_Rh103', 'score_name': 'reaction_Rh103_l10surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('RH103_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 0, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (11, 20), 'score_index': 0, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 16594.9,
error: 367.99804761300004,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 585.8918,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 16594.9,
error: 367.99804761300004,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}}
Pour manipuler plus aisément les résultats un Browser est disponible :
[12]:
t4b = t4pres.to_browser()
[13]:
print(t4b)
Browser object -> Number of content items: 18, data key: 'results', available metadata keys: ['composition', 'concentration', 'energy_split_name', 'index', 'particle', 'reaction', 'reaction_on_nucleus', 'response_function', 'response_index', 'response_name', 'response_type', 'score_index', 'score_name', 'scoring_mode', 'scoring_zone_id', 'scoring_zone_type']
-> Number of globals: 5
Il permet notamment de faire la sélection des résultats à partir des métadonnées du cas considéré. Leur nom est donné dans le print du Browser.
Les résultats de Tripoli-4 apparaissent sous la forme d’une liste de réponses (ou d’une liste de scores) dans le listing de sortie. Le Browser permet de les sélectionner de manière plus efficace.
Les résultats sont encapsulés dans un Dataset.
Étape 2 : sélection des données grâce au Browser
On différencie dans chaque “réponse” les données, soit les résultats du calcul effectué, et les métadonnées qui permettent de l’identifier. Chaque réponse est un dictionnaire dont la clef 'results' correspond aux données. Le Browser est construit à partir de cela.
Le Browser contient notamment :
la liste des résultats et de leurs métadonnées (
content)un index sur ces résultats (
index), basé sur les métadonnéesles variables globales du batch (
globals)
À noter : il est possible d’utiliser une autre clef que 'results' pour les données, à condition de le spécifier à la créaction du Browser grâce à l’argument data_key.
Explorer le Browser
2 niveaux d’impression sont également disponibles pour le Browser.
Méthodes d’aide pour découvrir le Browser :
keys: pour obtenir toutes les clefs de l’index, soit le nom / identifiant des métadonnées du listing concernéavailable_values: pour obtenir les valeurs correspondant à ces métadonnées
Dans ces deux cas il s’agit de générateurs. Pour afficher correctement les résultats il faut les encapsuler dans une liste par exemple.
[14]:
print('{!s}'.format(t4b)) # équivalent à print(t4rb)
Browser object -> Number of content items: 18, data key: 'results', available metadata keys: ['composition', 'concentration', 'energy_split_name', 'index', 'particle', 'reaction', 'reaction_on_nucleus', 'response_function', 'response_index', 'response_name', 'response_type', 'score_index', 'score_name', 'scoring_mode', 'scoring_zone_id', 'scoring_zone_type']
-> Number of globals: 5
[15]:
print('{!r}'.format(t4b))
<class 'valjean.eponine.browser.Browser'>, (Content items: [{'response_function': 'REACTION', 'response_name': 'reaction_Rh103', 'score_name': 'reaction_Rh103_l10surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('RH103_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 0, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (11, 20), 'score_index': 0, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 16594.9,
error: 367.99804761300004,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 585.8918,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 16594.9,
error: 367.99804761300004,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 0}, {'response_function': 'REACTION', 'response_name': 'reaction_Rh103', 'score_name': 'reaction_Rh103_l10surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('RH103_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 0, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (12, 21), 'score_index': 1, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 3191.064,
error: 90.06005902512001,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 112.6622,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 3191.064,
error: 90.06005902512001,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 1}, {'response_function': 'REACTION', 'response_name': 'reaction_Rh103', 'score_name': 'reaction_Rh103_l10surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('RH103_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 0, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (12, 22), 'score_index': 2, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 1203.617,
error: 35.162358712469995,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 42.49436,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 1203.617,
error: 35.162358712469995,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 2}, {'response_function': 'REACTION', 'response_name': 'reaction_Rh103', 'score_name': 'reaction_Rh103_l10surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('RH103_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 0, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (12, 23), 'score_index': 3, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 1047.175,
error: 30.108333712999997,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 36.97106,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 1047.175,
error: 30.108333712999997,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 3}, {'response_function': 'REACTION', 'response_name': 'reaction_Rh103', 'score_name': 'reaction_Rh103_l10surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('RH103_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 0, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (12, 24), 'score_index': 4, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 397.2467,
error: 15.013601366805002,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 14.02501,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 397.2467,
error: 15.013601366805002,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 4}, {'response_function': 'REACTION', 'response_name': 'reaction_Rh103', 'score_name': 'reaction_Rh103_l10surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('RH103_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 0, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (12, 25), 'score_index': 5, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 98.96879,
error: 4.0786265571996,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 3.494146,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 98.96879,
error: 4.0786265571996,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 5}, {'response_function': 'REACTION', 'response_name': 'reaction_Rh103', 'score_name': 'reaction_Rh103_l10surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('RH103_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 0, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (12, 26), 'score_index': 6, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 40.50244,
error: 1.9534910047136,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 1.429961,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 40.50244,
error: 1.9534910047136,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 6}, {'response_function': 'REACTION', 'response_name': 'reaction_Rh103', 'score_name': 'reaction_Rh103_l10surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('RH103_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 0, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (14, 27), 'score_index': 7, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 18.79975,
error: 1.1705560938875,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.6637354,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 18.79975,
error: 1.1705560938875,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 7}, {'response_function': 'REACTION', 'response_name': 'reaction_Rh103', 'score_name': 'reaction_Rh103_l10surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('RH103_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 0, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (14, 28), 'score_index': 8, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 5.299455,
error: 0.3073416807468,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.1871001,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 5.299455,
error: 0.3073416807468,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 8}, {'response_function': 'REACTION', 'response_name': 'reaction_Rh103', 'score_name': 'reaction_Rh103_l10surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('RH103_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 0, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (16, 29), 'score_index': 9, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 1.656025,
error: 0.08065141490524999,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.05846686,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 1.656025,
error: 0.08065141490524999,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 9}, {'response_function': 'REACTION', 'response_name': 'reaction_In115', 'score_name': 'reaction_In115_l3surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('IN115_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 1, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (14, 27), 'score_index': 0, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 3.279737,
error: 0.16236711908518,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.1157929,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 3.279737,
error: 0.16236711908518,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 10}, {'response_function': 'REACTION', 'response_name': 'reaction_In115', 'score_name': 'reaction_In115_l3surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('IN115_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 1, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (14, 28), 'score_index': 1, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.7353725,
error: 0.050511670734875,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.02596272,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.7353725,
error: 0.050511670734875,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 11}, {'response_function': 'REACTION', 'response_name': 'reaction_In115', 'score_name': 'reaction_In115_l3surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('IN115_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 1, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (16, 29), 'score_index': 2, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.2086517,
error: 0.010065887983318,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.007366562,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.2086517,
error: 0.010065887983318,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 12}, {'response_function': 'REACTION', 'response_name': 'reaction_S32', 'score_name': 'reaction_S32_l3surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('S32_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 2, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (14, 27), 'score_index': 0, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 1.019999,
error: 0.07280398922347002,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.03601161,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 1.019999,
error: 0.07280398922347002,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 13}, {'response_function': 'REACTION', 'response_name': 'reaction_S32', 'score_name': 'reaction_S32_l3surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('S32_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 2, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (14, 28), 'score_index': 1, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.1516089,
error: 0.01610849110767,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.005352634,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.1516089,
error: 0.01610849110767,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 14}, {'response_function': 'REACTION', 'response_name': 'reaction_S32', 'score_name': 'reaction_S32_l3surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('S32_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 2, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (16, 29), 'score_index': 2, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.03666873,
error: 0.0029075714077661996,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.001294609,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.03666873,
error: 0.0029075714077661996,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 15}, {'response_function': 'FLUX', 'response_name': 'flux', 'score_name': 'flux_l2surf', 'energy_split_name': 'DEC_SPECTRE', 'particle': 'NEUTRON', 'response_type': 'score', 'response_index': 3, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (14, 27), 'score_index': 0, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 44, 1, 1, 1)
value: [0.1528255 0.1498575 0.4623927 0.4774755 0.5853392 0.4022689
0.5847542 0.5541887 1.329951 0.4184096 1.270775 0.8474643
2.227431 0.7243235 0.7575581 1.973943 4.411218 1.044785
1.759198 3.106783 2.932957 3.357995 2.410037 1.634667
2.14932 2.402607 1.557616 1.298836 1.619081 1.204458
0.9177346 0.7668855 0.8060953 0.7114108 0.2920371 0.4349422
0.380331 0.3279657 0.3124642 0.3140535 0.1289041 0.1411806
0.08703017 0.08508481],
error: [0.05632055 0.05018791 0.12369583 0.11418201 0.19987718 0.11761779
0.1277354 0.1470686 0.2948367 0.09983287 0.26088197 0.42792337
1.05125321 0.25577992 0.22138613 0.75998345 1.58046045 0.22374113
0.27835948 1.17676365 0.49418419 0.85799627 0.57635312 0.3012451
0.43288272 0.55326417 0.21334152 0.17371282 0.19080271 0.13713802
0.12052911 0.11073857 0.10772327 0.09604771 0.05286599 0.08091682
0.05768724 0.05819502 0.0658838 0.08481524 0.03489702 0.05760408
0.03272591 0.04466015],
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([ 0.0463, 0.0525, 0.0595, 0.0674, 0.0764, 0.0865, 0.097 ,
0.111 , 0.126 , 0.143 , 0.162 , 0.183 , 0.207 , 0.235 ,
0.266 , 0.302 , 0.342 , 0.388 , 0.439 , 0.498 , 0.564 ,
0.639 , 0.724 , 0.821 , 0.93 , 1.054 , 1.194 , 1.353 ,
1.534 , 1.738 , 1.969 , 2.231 , 2.528 , 2.865 , 3.246 ,
3.679 , 4.169 , 4.724 , 5.353 , 6.065 , 6.873 , 7.788 ,
8.825 , 10. , 20. ])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'flux', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 44, 1, 1, 1)
value: [ 1.216075 1.197297 3.708972 3.809515 4.714329 3.511222
4.337321 4.372238 10.50823 3.35394 10.42561 6.876946
17.55725 5.845523 5.968283 15.86978 34.95567 8.460204
13.95075 24.96327 23.49181 26.88829 19.16809 13.11284
17.17214 19.26454 12.45939 10.34481 12.96755 9.651824
7.346338 6.136133 6.441557 5.69789 2.33224 3.478555
3.043143 2.623699 2.502168 2.511099 1.03137 1.129404
0.6962608 0.1227514],
error: [nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan
nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan
nan nan nan nan nan nan nan nan],
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([ 0.0463, 0.0525, 0.0595, 0.0674, 0.0764, 0.0865, 0.097 ,
0.111 , 0.126 , 0.143 , 0.162 , 0.183 , 0.207 , 0.235 ,
0.266 , 0.302 , 0.342 , 0.388 , 0.439 , 0.498 , 0.564 ,
0.639 , 0.724 , 0.821 , 0.93 , 1.054 , 1.194 , 1.353 ,
1.534 , 1.738 , 1.969 , 2.231 , 2.528 , 2.865 , 3.246 ,
3.679 , 4.169 , 4.724 , 5.353 , 6.065 , 6.873 , 7.788 ,
8.825 , 10. , 20. ])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'flux/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 49.51463,
error: 3.5046068899886,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([ 0.0463, 20. ])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'flux'}, 'index': 16}, {'response_function': 'FLUX', 'response_name': 'flux', 'score_name': 'flux_l2surf', 'energy_split_name': 'DEC_SPECTRE', 'particle': 'NEUTRON', 'response_type': 'score', 'response_index': 3, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (16, 29), 'score_index': 1, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 44, 1, 1, 1)
value: [0.03050079 0.06832957 0.07695755 0.09925667 0.1350992 0.02155418
0.08208997 0.1378661 0.3047329 0.07209889 0.2593902 0.1593264
0.2394927 0.1843582 0.4052755 0.3252563 0.4268424 0.1484201
0.2554633 0.2897066 0.4703281 0.498303 0.2471051 0.2301476
0.2617021 0.1997403 0.1476602 0.121049 0.09662921 0.09268166
0.05543372 0.04428578 0.03727038 0.02329219 0.01471019 0.01064267
0.01057807 0.010661 0.00989727 0.00583138 0.00626957 0.00349015
0.00309229 0.00675048],
error: [0.01210843 0.03119505 0.01720868 0.01918197 0.02985159 0.00884732
0.01680104 0.02190428 0.03902812 0.01818942 0.03283141 0.03104628
0.09498242 0.06235245 0.09121256 0.059099 0.08275433 0.0348335
0.04874998 0.05476054 0.07168115 0.07391498 0.0579394 0.04094811
0.06295158 0.02666253 0.02132944 0.01697825 0.01354986 0.01089185
0.00827571 0.00729076 0.00627467 0.00498183 0.00346623 0.00270574
0.00260966 0.0023411 0.00240927 0.00189545 0.00196971 0.00159711
0.00149497 0.0047856 ],
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([ 0.0463, 0.0525, 0.0595, 0.0674, 0.0764, 0.0865, 0.097 ,
0.111 , 0.126 , 0.143 , 0.162 , 0.183 , 0.207 , 0.235 ,
0.266 , 0.302 , 0.342 , 0.388 , 0.439 , 0.498 , 0.564 ,
0.639 , 0.724 , 0.821 , 0.93 , 1.054 , 1.194 , 1.353 ,
1.534 , 1.738 , 1.969 , 2.231 , 2.528 , 2.865 , 3.246 ,
3.679 , 4.169 , 4.724 , 5.353 , 6.065 , 6.873 , 7.788 ,
8.825 , 10. , 20. ])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'flux', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 44, 1, 1, 1)
value: [0.2427032 0.5459238 0.6172965 0.7919144 1.088091 0.1881366
0.6088893 1.087686 2.407761 0.5779393 2.128071 1.292891
1.88775 1.48783 3.192889 2.614941 3.382414 1.20184
2.025869 2.327817 3.76714 3.990034 1.965336 1.84618
2.090888 1.601554 1.181136 0.9641162 0.7739233 0.7426966
0.4437392 0.3543468 0.2978298 0.1865537 0.1174771 0.08511736
0.08463831 0.08528712 0.07925593 0.04662634 0.05016322 0.02792023
0.024739 0.00973889],
error: [nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan
nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan nan
nan nan nan nan nan nan nan nan],
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([ 0.0463, 0.0525, 0.0595, 0.0674, 0.0764, 0.0865, 0.097 ,
0.111 , 0.126 , 0.143 , 0.162 , 0.183 , 0.207 , 0.235 ,
0.266 , 0.302 , 0.342 , 0.388 , 0.439 , 0.498 , 0.564 ,
0.639 , 0.724 , 0.821 , 0.93 , 1.054 , 1.194 , 1.353 ,
1.534 , 1.738 , 1.969 , 2.231 , 2.528 , 2.865 , 3.246 ,
3.679 , 4.169 , 4.724 , 5.353 , 6.065 , 6.873 , 7.788 ,
8.825 , 10. , 20. ])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'flux/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 6.329569,
error: 0.32567252874664,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([ 0.0463, 20. ])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'flux'}, 'index': 17}], Index: defaultdict(<function _make_defaultdict_set at 0x7efe6c30e700>, {'response_function': defaultdict(<class 'set'>, {'REACTION': {0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15}, 'FLUX': {16, 17}}), 'response_name': defaultdict(<class 'set'>, {'reaction_Rh103': {0, 1, 2, 3, 4, 5, 6, 7, 8, 9}, 'reaction_In115': {10, 11, 12}, 'reaction_S32': {13, 14, 15}, 'flux': {16, 17}}), 'score_name': defaultdict(<class 'set'>, {'reaction_Rh103_l10surf': {0, 1, 2, 3, 4, 5, 6, 7, 8, 9}, 'reaction_In115_l3surf': {10, 11, 12}, 'reaction_S32_l3surf': {13, 14, 15}, 'flux_l2surf': {16, 17}}), 'energy_split_name': defaultdict(<class 'set'>, {'DEC_INTEGRAL': {0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15}, 'DEC_SPECTRE': {16, 17}}), 'particle': defaultdict(<class 'set'>, {'NEUTRON': {0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17}}), 'response_type': defaultdict(<class 'set'>, {'score': {0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17}}), 'reaction_on_nucleus': defaultdict(<class 'set'>, {('RH103_IRDF85',): {0, 1, 2, 3, 4, 5, 6, 7, 8, 9}, ('IN115_IRDF85',): {10, 11, 12}, ('S32_IRDF85',): {13, 14, 15}}), 'composition': defaultdict(<class 'set'>, {('none',): {0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15}}), 'concentration': defaultdict(<class 'set'>, {(1.0,): {0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15}}), 'reaction': defaultdict(<class 'set'>, {('tabulated data',): {0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15}}), 'response_index': defaultdict(<class 'set'>, {0: {0, 1, 2, 3, 4, 5, 6, 7, 8, 9}, 1: {10, 11, 12}, 2: {13, 14, 15}, 3: {16, 17}}), 'scoring_mode': defaultdict(<class 'set'>, {'SCORE_SURF': {0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17}}), 'scoring_zone_type': defaultdict(<class 'set'>, {'Frontier': {0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17}}), 'scoring_zone_id': defaultdict(<class 'set'>, {(11, 20): {0}, (12, 21): {1}, (12, 22): {2}, (12, 23): {3}, (12, 24): {4}, (12, 25): {5}, (12, 26): {6}, (14, 27): {16, 10, 13, 7}, (14, 28): {8, 11, 14}, (16, 29): {9, 12, 17, 15}}), 'score_index': defaultdict(<class 'set'>, {0: {0, 16, 10, 13}, 1: {1, 11, 14, 17}, 2: {2, 12, 15}, 3: {3}, 4: {4}, 5: {5}, 6: {6}, 7: {7}, 8: {8}, 9: {9}}), 'index': defaultdict(<class 'set'>, {0: {0}, 1: {1}, 2: {2}, 3: {3}, 4: {4}, 5: {5}, 6: {6}, 7: {7}, 8: {8}, 9: {9}, 10: {10}, 11: {11}, 12: {12}, 13: {13}, 14: {14}, 15: {15}, 16: {16}, 17: {17}})}))
[16]:
list(t4b.keys())
[16]:
['response_function',
'response_name',
'score_name',
'energy_split_name',
'particle',
'response_type',
'reaction_on_nucleus',
'composition',
'concentration',
'reaction',
'response_index',
'scoring_mode',
'scoring_zone_type',
'scoring_zone_id',
'score_index',
'index']
[17]:
print('values for score_name:', list(t4b.available_values('score_name')))
print('values for response_function:', list(t4b.available_values('response_function')))
print('values for scoring_zone_id:', list(t4b.available_values('scoring_zone_id')))
print('values for particle:', list(t4b.available_values('particle')))
print('values for reaction_on_nucleus:', list(t4b.available_values('reaction_on_nucleus')))
values for score_name: ['reaction_Rh103_l10surf', 'reaction_In115_l3surf', 'reaction_S32_l3surf', 'flux_l2surf']
values for response_function: ['REACTION', 'FLUX']
values for scoring_zone_id: [(11, 20), (12, 21), (12, 22), (12, 23), (12, 24), (12, 25), (12, 26), (14, 27), (14, 28), (16, 29)]
values for particle: ['NEUTRON']
values for reaction_on_nucleus: [('RH103_IRDF85',), ('IN115_IRDF85',), ('S32_IRDF85',)]
Sélection des réponses / résultats grâce au Browser
Il est possible de sélectionner les réponses à partir des métadonnées. La méthode à choisir dépend de l’objet dont on a besoin en sortie.
filter_bypour récupérer un sous-Browserselect_bypour récupérer la réponse si elle est unique
La sélection se fait
par mot-clef/valeur, soit métadonnée/valeur de cette métadonnée (keyword arguments correspondant aux clefs disponibles)
include=tuple(key)pour sélectionner toutes les réponses contenant cette métadonnée sans distinction de la valeurexclude=tuple(key)pour exclure les réponses contenant une métadonnée
include et exclude ne fonctionnent que sur le nom de la métadonnée, pas sur sa valeur.
Ces 3 possibilités sont bien sûr combinables.
[18]:
b_flux = t4b.filter_by(response_function='FLUX')
print(b_flux)
Browser object -> Number of content items: 2, data key: 'results', available metadata keys: ['energy_split_name', 'index', 'particle', 'response_function', 'response_index', 'response_name', 'response_type', 'score_index', 'score_name', 'scoring_mode', 'scoring_zone_id', 'scoring_zone_type']
-> Number of globals: 5
Le nombre de réponses dans le Browser n’est maintenant plus que de 2 au lieu de 18, il ne reste plus que celles correspondant à un flux. Récupérer les réponses peut se faire directement ou avec l’autre méthode.
Le sous-Browser remet à zéro les index (en crée un nouveau en réalité).
À noter : les variables globales du Browser sont transmises au sous-Browser.
[19]:
b_flux.globals == t4b.globals
[19]:
True
[20]:
b_excl_compo = t4b.filter_by(exclude=('composition',)) # attention à bien utiliser un tuple
print(b_excl_compo)
Browser object -> Number of content items: 2, data key: 'results', available metadata keys: ['energy_split_name', 'index', 'particle', 'response_function', 'response_index', 'response_name', 'response_type', 'score_index', 'score_name', 'scoring_mode', 'scoring_zone_id', 'scoring_zone_type']
-> Number of globals: 5
Les réponses ont été « aplaties » (ou flattened). Dans le jeu de données d’origine 3 réponses sont demandées sur une liste de scoring zones à chaque fois.
SCORE
4
NAME reaction_Rh103_l10surf
reaction_Rh103
SURF DECOUPAGE DEC_INTEGRAL FRONTIER LIST 10
11 20 12 21 12 22 12 23 12 24
12 25 12 26 14 27 14 28 16 29
NAME reaction_In115_l3surf
reaction_In115
SURF DECOUPAGE DEC_INTEGRAL FRONTIER LIST 3
14 27 14 28 16 29
NAME reaction_S32_l3surf
reaction_S32
SURF DECOUPAGE DEC_INTEGRAL FRONTIER LIST 3
14 27 14 28 16 29
NAME flux_l2surf
flux
SURF DECOUPAGE DEC_SPECTRE FRONTIER LIST 2
14 27 16 29
FIN_SCORE
Pour récupérer une réponse donnée dans le cas présent il faut au moins deux critères, dont scoring_zone_id.
[21]:
b_reacIn_16_29 = t4b.filter_by(score_name='reaction_In115_l3surf', scoring_zone_id=(16, 29))
print(b_reacIn_16_29)
Browser object -> Number of content items: 1, data key: 'results', available metadata keys: ['composition', 'concentration', 'energy_split_name', 'index', 'particle', 'reaction', 'reaction_on_nucleus', 'response_function', 'response_index', 'response_name', 'response_type', 'score_index', 'score_name', 'scoring_mode', 'scoring_zone_id', 'scoring_zone_type']
-> Number of globals: 5
Sélection d’une réponse donnée : select_by
[22]:
reacIn_16_29 = t4b.select_by(score_name='reaction_In115_l3surf', scoring_zone_id=(16, 29))
print(type(reacIn_16_29))
print(reacIn_16_29)
<class 'dict'>
{'response_function': 'REACTION', 'response_name': 'reaction_In115', 'score_name': 'reaction_In115_l3surf', 'energy_split_name': 'DEC_INTEGRAL', 'particle': 'NEUTRON', 'response_type': 'score', 'reaction_on_nucleus': ('IN115_IRDF85',), 'composition': ('none',), 'concentration': (1.0,), 'reaction': ('tabulated data',), 'response_index': 1, 'scoring_mode': 'SCORE_SURF', 'scoring_zone_type': 'Frontier', 'scoring_zone_id': (16, 29), 'score_index': 2, 'results': {'discarded_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 0.000000e+00, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'discarded_batches'
, 'used_batches': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.int64'>
value: 1.000000e+02, error: 0.000000e+00, bins: OrderedDict()
name: '', what: 'used_batches'
, 'score': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.2086517,
error: 0.010065887983318,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction', 'score/lethargy': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.007366562,
error: nan,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction/lethargy', 'units': {'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}, 'score_integrated': class: <class 'valjean.eponine.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape: (1, 1, 1, 1, 1, 1, 1)
value: 0.2086517,
error: 0.010065887983318,
bins: OrderedDict([('u', array([], dtype=float64)), ('v', array([], dtype=float64)), ('w', array([], dtype=float64)), ('e', array([1.e-11, 2.e+01])), ('t', array([], dtype=float64)), ('mu', array([], dtype=float64)), ('phi', array([], dtype=float64))]),
name: '', what: 'reaction'}, 'index': 12}
[23]:
flux_14_27 = t4b.select_by(response_function='FLUX', scoring_zone_id=(14, 27))
print(type(flux_14_27))
print(list(flux_14_27['results'].keys()))
<class 'dict'>
['discarded_batches', 'used_batches', 'score', 'score/lethargy', 'units', 'score_integrated']
Une fois le résultat voulu sélectionné on peut accéder aux données elles-mêmes.
Dans le cas de flux_14_27 (flux surfacique entre les surfaces 14 et 27) il existe 6 types de données :
les nombres de batches utilisés et mis de côté pour le calcul du flux
le spectre utilisant tout le découpage en énergie (score demandé) (
score)l’intégrale du spectre sur le découpage en énergie (
score_integrated)le spectre score/lethargy, soit renormalisé à la largeur du bin, sur tout le découpage en énergie (
score/lethargy)les unités
Tous ces résultats sont encapsulés dans un Dataset, à l’exception des unités qui sont dans un dictionnaire.
Étape 3 : utilisation d’un Dataset
Le Dataset permet de comparer les données à d’autres grâce aux tests ou de les représenter par exemple.
Le Dataset permet de stocker les données et de faire des opérations dessus (ajout de datasets, sélection de dimension, multiplication par un dataset ou une constante, etc.). Les tests statistiques proposés par valjean attendent également des Dataset. Leur argument name est mis par défaut à celui du fichier parsé. L’argument what est utilisé pour donner à un nom au type de données qu’ils contiennent. Il sera utilisé en axe des ordonnées d’un histogramme 1D par exemple. Ces
deux arguments sont modifiables à tout moment.
[24]:
fds_flux_14_27 = flux_14_27['results']['score']
print(fds_flux_14_27)
fds_flux_14_27.name = 'flux(14, 27)'
fds_flux_14_27.what = 'Flux'
shape: (1, 1, 1, 44, 1, 1, 1), dim: 7, type: <class 'numpy.ndarray'>, bins: ['u: []', 'v: []', 'w: []', 'e: [ 0.0463 0.0525 0.0595 0.0674 0.0764 0.0865 0.097 0.111 0.126 0.143 0.162 0.183 0.207 0.235 0.266 0.302 0.342 0.388 0.439 0.498 0.564 0.639 0.724 0.821 0.93 1.054 1.194 1.353 1.534 1.738 1.969 2.231 2.528 2.865 3.246 3.679 4.169 4.724 5.353 6.065 6.873 7.788 8.825 10. 20. ]', 't: []', 'mu: []', 'phi: []'], name: , what: flux
Les spectres (comme les maillages) sont par défault en 7 dimensions, dont les noms sont stockés dans les bins. Le dictionnaire de bins est un OrderedDict qui respecte l’ordre des dimensions de l’array Numpy.
Petit commentaire sur les noms de ces bins :
'u','v','w'correspondent aux 3 variables d’espace (surtout pour les maillages), ce qui peut donc être par exemple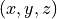 ,
,  ou
ou 
'e'l’énergie,'t'le temps'mu'et'phi'la direction de la particule
Des unités sont données par défaut (celle de Tripoli par défaut si non précisé, unknown pour les variables d’espace).
[25]:
print(flux_14_27['results']['units'])
{'u': 'cm', 'v': 'unknown', 'w': 'unknown', 'e': 'MeV', 't': 's', 'mu': '', 'phi': 'rad', 'score': 'unknown', 'sigma': '%'}
La plupart du temps beaucoup de ces dimensions ne sont pas précisées dans Tripoli-4, n’ont donc pas de découpage (ou de bins) et sont ainsi à 1 dans la shape. Pour les réduire, pour l’affichage ou l’utilisation, il est possible de squeezer le Dataset. Cette suppression des dimensions non utilisées en conseillée pour les tests.
Dans le cas de flux_14_27 seule la dimension correspondant à l’énergie est pertinente, le squeeze permet de ne conserver qu’elle.
[26]:
ds_flux_14_27 = fds_flux_14_27.squeeze()
print(ds_flux_14_27)
shape: (44,), dim: 1, type: <class 'numpy.ndarray'>, bins: ['e: [ 0.0463 0.0525 0.0595 0.0674 0.0764 0.0865 0.097 0.111 0.126 0.143 0.162 0.183 0.207 0.235 0.266 0.302 0.342 0.388 0.439 0.498 0.564 0.639 0.724 0.821 0.93 1.054 1.194 1.353 1.534 1.738 1.969 2.231 2.528 2.865 3.246 3.679 4.169 4.724 5.353 6.065 6.873 7.788 8.825 10. 20. ]'], name: flux(14, 27), what: Flux
Le Dataset correspondant au résultat intégré du spectre donne quant à lui :
[27]:
ds_flux_14_27_int = flux_14_27['results']['score_integrated']
print(ds_flux_14_27_int)
ds_flux_14_27_int.name='flux_14_27'
ds_flux_14_27_int.what='Flux'
print(ds_flux_14_27_int.squeeze())
shape: (1, 1, 1, 1, 1, 1, 1), dim: 7, type: <class 'numpy.ndarray'>, bins: ['u: []', 'v: []', 'w: []', 'e: [ 0.0463 20. ]', 't: []', 'mu: []', 'phi: []'], name: , what: flux
shape: (), dim: 0, type: <class 'numpy.ndarray'>, bins: [], name: flux_14_27, what: Flux
Le squeeze supprime les bins dans deux conditions :
il n’y a pas de bins
il n’y a qu’un bin, donc la dimension est triviale
Remarque : pour une dimension donnée, si on a N valeurs, les bins sont
soit donnés par leurs limites, il y a donc N+1 valeurs de bins,
soit donnés par leurs centres, il y a donc N valeurs
Pour construire un Dataset on doit cependant connaître les clefs disponibles sous results.
Dans le cas des taux de réaction on a :
[28]:
print(list(reacIn_16_29['results'].keys()))
['discarded_batches', 'used_batches', 'score', 'score/lethargy', 'units', 'score_integrated']
Dans ce cas précis, comme le découpage ne comprend qu’un seul bin, le Dataset issu de 'integrated' et celui issu de 'spectrum' devraient être les mêmes.
[29]:
ds_reacIn_16_29_spec = reacIn_16_29['results']['score']
ds_reacIn_16_29_spec.name='spectrum'
ds_reacIn_16_29_spec.what='Flux'
print(ds_reacIn_16_29_spec)
print(ds_reacIn_16_29_spec.value)
ds_reacIn_16_29_int = reacIn_16_29['results']['score_integrated']
ds_reacIn_16_29_int.name='integrated'
ds_reacIn_16_29_int.what='Flux'
print(ds_reacIn_16_29_int)
print(ds_reacIn_16_29_int.value)
shape: (1, 1, 1, 1, 1, 1, 1), dim: 7, type: <class 'numpy.ndarray'>, bins: ['u: []', 'v: []', 'w: []', 'e: [1.e-11 2.e+01]', 't: []', 'mu: []', 'phi: []'], name: spectrum, what: Flux
[[[[[[[0.2086517]]]]]]]
shape: (1, 1, 1, 1, 1, 1, 1), dim: 7, type: <class 'numpy.ndarray'>, bins: ['u: []', 'v: []', 'w: []', 'e: [1.e-11 2.e+01]', 't: []', 'mu: []', 'phi: []'], name: integrated, what: Flux
[[[[[[[0.2086517]]]]]]]
Étape 4 : faire un test et le représenter, exemple du test de Student
Pour avoir une description du test de Student, voir la documentation de valjean.
[30]:
from valjean.gavroche.stat_tests.student import TestStudent
La représentation d’un test peut se faire sous forme de tableau ou de graphique.
[31]:
# includes et initialisations pour les tableaux
from valjean.javert.representation import TableRepresenter
from valjean.javert.rst import RstFormatter
tabrepr = TableRepresenter()
rstformat = RstFormatter()
[32]:
# include et initialisation pour les graphiques
from valjean.javert.representation import PlotRepresenter
from valjean.javert.mpl import MplPlot
plotrepr = PlotRepresenter()
Comparaison de l’intégrale du spectre et du spectre dans le cas de la réaction sur In115 entre les surfaces 16 et 29
[33]:
stud_reacIn = TestStudent(ds_reacIn_16_29_spec, ds_reacIn_16_29_int, name='spectrum vs intregral',
description="Comparaison du spectre sur 1 bin à l'intégrale").evaluate()
print(type(stud_reacIn))
print(bool(stud_reacIn))
<class 'valjean.gavroche.stat_tests.student.TestResultStudent'>
True
[34]:
montab = tabrepr(stud_reacIn) # il s'agit d'une liste de TableTemplate
monrst = rstformat.template(montab[0])
print(monrst)
Student test: OK
[35]:
monplot = plotrepr(stud_reacIn)
print(monplot)
[]
Les graphiques ne sont pas disponibles pour les quantités sans réels bins.
Comparaison des flux entre les surfaces 14 et 27 et les surfaces 16 et 29
Le découpage en énergie de ces deux flux étant les mêmes leur comparaison est possible.
[36]:
# construction du Dataset pour le flux entre les surfaces 16 et 29
flux_16_29 = t4b.select_by(response_function='FLUX', scoring_zone_id=(16, 29))
ds_flux_16_29 = flux_16_29['results']['score'].squeeze()
ds_flux_16_29.name='flux(16, 29)'
ds_flux_16_29.what='Flux'
[37]:
tstud_flux = TestStudent(ds_flux_14_27, ds_flux_16_29, name="flux_14_17_vs_16_29",
description="Comparison of the flux between surfaces 14 and 27 and surfaces 16 and 29")
stud_flux = tstud_flux.evaluate()
print(bool(stud_flux))
False
[38]:
montab = tabrepr(stud_flux) # encore une liste
monrst = rstformat.template(montab[0])
print(monrst)
.. role:: hl
.. table::
:widths: auto
=============== =============== =============== =============== =============== =========== ===========
e v(flux(14, 27)) σ(flux(14, 27)) v(flux(16, 29)) σ(flux(16, 29)) t Student?
=============== =============== =============== =============== =============== =========== ===========
0.0595 - 0.0674 0.462393 0.123696 0.0769575 0.0172087 3.08627 :hl:`False`
0.0674 - 0.0764 0.477475 0.114182 0.0992567 0.019182 3.26665 :hl:`False`
0.0865 - 0.097 0.402269 0.117618 0.0215542 0.00884732 3.22776 :hl:`False`
0.097 - 0.111 0.584754 0.127735 0.08209 0.016801 3.90159 :hl:`False`
0.111 - 0.126 0.554189 0.147069 0.137866 0.0219043 2.79992 :hl:`False`
0.126 - 0.143 1.32995 0.294837 0.304733 0.0390281 3.44717 :hl:`False`
0.143 - 0.162 0.41841 0.0998329 0.0720989 0.0181894 3.41272 :hl:`False`
0.162 - 0.183 1.27077 0.260882 0.25939 0.0328314 3.84645 :hl:`False`
0.388 - 0.439 1.04479 0.223741 0.14842 0.0348335 3.95857 :hl:`False`
0.439 - 0.498 1.7592 0.278359 0.255463 0.04875 5.32114 :hl:`False`
0.564 - 0.639 2.93296 0.494184 0.470328 0.0716812 4.93161 :hl:`False`
0.639 - 0.724 3.35799 0.857996 0.498303 0.073915 3.32069 :hl:`False`
0.724 - 0.821 2.41004 0.576353 0.247105 0.0579394 3.73397 :hl:`False`
0.821 - 0.93 1.63467 0.301245 0.230148 0.0409481 4.6199 :hl:`False`
0.93 - 1.054 2.14932 0.432883 0.261702 0.0629516 4.31519 :hl:`False`
1.054 - 1.194 2.40261 0.553264 0.19974 0.0266625 3.97697 :hl:`False`
1.194 - 1.353 1.55762 0.213342 0.14766 0.0213294 6.57613 :hl:`False`
1.353 - 1.534 1.29884 0.173713 0.121049 0.0169782 6.74793 :hl:`False`
1.534 - 1.738 1.61908 0.190803 0.0966292 0.0135499 7.95915 :hl:`False`
1.738 - 1.969 1.20446 0.137138 0.0926817 0.0108918 8.08154 :hl:`False`
1.969 - 2.231 0.917735 0.120529 0.0554337 0.00827571 7.13749 :hl:`False`
2.231 - 2.528 0.766885 0.110739 0.0442858 0.00729076 6.51118 :hl:`False`
2.528 - 2.865 0.806095 0.107723 0.0372704 0.00627467 7.12496 :hl:`False`
2.865 - 3.246 0.711411 0.0960477 0.0232922 0.00498183 7.15472 :hl:`False`
3.246 - 3.679 0.292037 0.052866 0.0147102 0.00346623 5.23461 :hl:`False`
3.679 - 4.169 0.434942 0.0809168 0.0106427 0.00270574 5.24072 :hl:`False`
4.169 - 4.724 0.380331 0.0576872 0.0105781 0.00260966 6.40307 :hl:`False`
4.724 - 5.353 0.327966 0.058195 0.010661 0.0023411 5.44803 :hl:`False`
5.353 - 6.065 0.312464 0.0658838 0.00989727 0.00240927 4.58937 :hl:`False`
6.065 - 6.873 0.314053 0.0848152 0.00583138 0.00189545 3.63313 :hl:`False`
6.873 - 7.788 0.128904 0.034897 0.00626957 0.00196971 3.5086 :hl:`False`
=============== =============== =============== =============== =============== =========== ===========
[39]:
monplot = plotrepr(stud_flux)
mpl = MplPlot(monplot[0]).draw()

Dans le cas d’un spectre un test de Student est effectué par bin. Dans le cas présent les spectres ne sont manifestement pas en accord, ce qui est attendu. Cependant, dans le cas d’une comparaison avec des données réellement comparables on peut s’attendre à un certain nombre de bins pour lesquels la comparaison échoue mais pas tous. Ce nombre dépend du nombre de bins.
Pour évaluer cela un autre test statistique est possible, le test de Holm-Bonferroni (voir la documentation pour plus de précisions).