dataset – Unified data format
Common module to structure data in same way for all codes.
Dataset definition and initialisation
A dataset is composed by 5 members:
value: a
numpy.ndarrayor anumpy.generic(scalar from numpy represented like the arrays, with dim, etc)error: an object of same type as value
bins: an
collections.OrderedDict(optional and named argument)name: name of the dataset (optional and named argument)
what: can be used to store the name of the quantity represented by the dataset (optional and named argument)
The bins object should have the same dimension as the value, the order matches
the dimensions. If no bins are available it is still possible to use an empty
collections.OrderedDict.
>>> from valjean.eponine.dataset import Dataset
>>> import numpy as np
>>> from collections import OrderedDict
>>> bins = OrderedDict([('e', np.array([1, 2, 3])), ('t', np.arange(5))])
>>> ds1 = Dataset(np.arange(10).reshape(2, 5),
... np.array([0.3]*10).reshape(2, 5),
... bins=bins, name='ds1', what='spam')
>>> ds1.name
'ds1'
>>> ds1.what
'spam'
>>> len(bins) == ds1.ndim
True
>>> ds1.error.shape == ds1.value.shape
True
>>> ds1.value.shape == (2, 5)
True
>>> np.array_equal(ds1.value, [[0, 1, 2, 3, 4], [5, 6, 7, 8, 9]])
True
>>> np.array_equal(ds1.error,
... [[0.3, 0.3, 0.3, 0.3, 0.3], [0.3, 0.3, 0.3, 0.3, 0.3]])
True
>>> list(bins.keys())
['e', 't']
>>> np.array_equal(bins['e'], [1, 2, 3])
True
>>> np.array_equal(bins['t'], [0, 1, 2, 3, 4])
True
A new dataset can also be created from an already existing one, using copy.
No matter how the dataset is generated, attributes can be changed afterwards:
>>> nds = ds1.copy()
>>> nds.name = 'egg'
>>> nds.what = ''
>>> print(f'name: ds1={ds1.name!r}, nds={nds.name!r}')
name: ds1='ds1', nds='egg'
>>> print(f'what: ds1={ds1.what!r}, nds={nds.what!r}')
what: ds1='spam', nds=''
>>> np.array_equal(ds1.value, nds.value)
True
Errors are emitted if the arguments do not have the expected type or if the shapes or dimensions are not consistent:
>>> tds = Dataset([1, 2, 3], [0.5, 0.5, 0.5])
Traceback (most recent call last):
[...]
TypeError: value does not have the expected type (numpy.ndarray or numpy.generic = scalar)
>>> tds = Dataset(np.arange(6).reshape(2, 3), np.arange(6).reshape(3, 2))
Traceback (most recent call last):
[...]
ValueError: Value and error do not have the same shape
>>> tds = Dataset(np.arange(6).reshape(2, 3),
... np.array([0.5]*6).reshape(2, 3),
... bins={'spam': [1, 2], 'egg': [1, 2, 3]})
Traceback (most recent call last):
[...]
TypeError: bins should be an OrderedDict
>>> tds = Dataset(np.arange(6).reshape(2, 3),
... np.array([0.5]*6).reshape(2, 3),
... bins=OrderedDict([('spam', [1, 2])]))
Traceback (most recent call last):
[...]
ValueError: Number of dimensions of bins does not correspond to number of dimensions of value
Squeezing a dataset
If there are useless dimensions, the dataset can be squeezed:
>>> vals = np.arange(6).reshape(1, 2, 1, 3)
>>> errs = np.array([0.1]*6).reshape(1, 2, 1, 3)
>>> bins2 = OrderedDict([('bacon', np.array([0, 1])),
... ('egg', np.array([0, 2, 4])),
... ('sausage', np.array([10, 20])),
... ('spam', np.array([-5, 0, 5, 10]))])
>>> ds = Dataset(vals, errs, bins=bins2)
>>> ds.value.shape == (1, 2, 1, 3)
True
>>> len(ds.bins) == 4
True
>>> np.array_equal(ds.value, np.array([[[[0, 1, 2]], [[3, 4, 5]]]]))
True
>>> sds = ds.squeeze()
>>> sds.ndim
2
>>> len(sds.bins) == 2
True
>>> sds.shape
(2, 3)
>>> list(len(x)-1 for x in sds.bins.values()) # edges of bins, so N+1
[2, 3]
>>> list(sds.bins.keys()) == ['egg', 'spam']
True
>>> np.array_equal(sds.value, np.array([[0, 1, 2], [3, 4, 5]]))
True
The dimensions with only one bin are squeezed, the same is done on the bins.
>>> nsds = ds.squeeze()
>>> np.array_equal(nsds.value, sds.value)
True
>>> nsds.bins == sds.bins
True
Operations available on Datasets
Standard operations, (+, -, *, /, []), are available for Dataset.
Examples of their use are given, including failing cases. Some are used in
various methods but shown only once.
All operations conserve the name of the initial dataset.
The what attribute can be used to store the name of the quantity represented by the dataset. It is updated if it involves another dataset:
in addition or substraction, if what is different it contains both separated by the symbol
in multiplication or division it always contains both separeted by the symbol
Addition and subtraction
Addition and subtraction are possible between a Dataset and a scalar
(numpy.generic), a numpy.ndarray and another Dataset.
The operator to use for addition is + and for subtraction -.
Restrictions in addition or subtraction with a numpy.ndarray are handled
by NumPy.
The addition or subtraction of two Dataset can be done if
both values have the same shape (
consistent_datasets)bins conditions (
same_coords):EITHER the second
Datasetdoes not have any binsOR bins are the same, i.e. have the same keys and bins values
Example addition of a scalar value (only on value)
>>> ds1p10 = ds1 + 10
>>> np.array_equal(ds1p10.value,
... [[10, 11, 12, 13, 14], [15, 16, 17, 18, 19]])
True
>>> np.array_equal(ds1p10.error, ds1.error)
True
>>> ds1p10.bins == ds1.bins
True
>>> ds1p10.what == ds1.what
True
As expected it ‘only’ acts on the value, error and bins are unchanged.
Example of subtraction of a numpy.ndarray
>>> a = np.array([100]*10).reshape(2, 5)
>>> ds1.value.shape == a.shape
True
>>> ds1ma = ds1 - a
>>> ds1ma.__class__
<class 'valjean.eponine.dataset.Dataset'>
>>> np.array_equal(ds1ma.value, [[-100, -99, -98, -97, -96],
... [-95, -94, -93, -92, -91]])
True
>>> np.array_equal(ds1ma.error, ds1.error)
True
>>> ds1ma.bins == ds1.bins
True
>>> ds1ma.what == ds1.what
True
a and ds1 have the same shape to everything is fine.
>>> b = np.array([100]*10)
>>> ds1.value.shape == b.shape
False
>>> ds1pb = ds1 + b
Traceback (most recent call last):
...
ValueError: operands could not be broadcast together with shapes (2,5) (10,)
If the shapes are not the same NumPy raises an exception (and ds1pb is not defined).
Example of addition or subtraction of another Dataset
>>> ds2 = Dataset(value=np.arange(20, 30).reshape(2, 5),
... error=np.array([0.4]*10).reshape(2, 5),
... bins=bins, name='ds2', what='spam')
>>> ds1.bins == ds2.bins
True
>>> ds1p2 = ds1 + ds2
>>> ds1p2.name == ds1.name
True
>>> ds1p2.what == ds1.what
True
>>> np.array_equal(ds1p2.value, [[20, 22, 24, 26, 28],
... [30, 32, 34, 36, 38]])
True
>>> np.array_equal(ds1p2.error, np.array([0.5]*10).reshape(2, 5))
True
The error is calulated considering both datasets are independent, so
quadratically (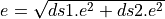 ).
).
Datasets without binning can also be added:
>>> ds3 = Dataset(value=np.arange(200, 300, 10).reshape(2, 5),
... error=np.array([0.4]*10).reshape(2, 5), name='ds3')
>>> ds3.bins
OrderedDict()
>>> ds1p3 = ds1 + ds3
>>> ds1p3.name == ds1.name
True
>>> ds1p3.what == ds1.what
False
>>> ds1p3.what
'spam+'
>>> np.array_equal(ds1p3.value, [[200, 211, 222, 233, 244],
... [255, 266, 277, 288, 299]])
True
>>> np.array_equal(ds1p3.error, np.array([0.5]*10).reshape(2, 5))
True
>>> same_coords(ds1p3, ds1)
True
Bins of the dataset on the left are kept.
Like in NumPy array addition, values need to have the same shape:
>>> ds4 = Dataset(np.arange(5), np.array([0.01]*5), name='ds4')
>>> f"shape ds1 {ds1.value.shape}, ds4 {ds4.value.shape} -> comp = {ds1.value.shape == ds4.value.shape}"
'shape ds1 (2, 5), ds4 (5,) -> comp = False'
>>> ds1 + ds4
Traceback (most recent call last):
[...]
ValueError: Datasets to add do not have same shape
If bins are given, they need to have the same keys and the same values.
>>> bins5 = OrderedDict([('E', np.array([1, 2, 3])), ('t', np.arange(5))])
>>> ds5 = Dataset(np.arange(0, -10, -1).reshape(2, 5),
... np.array([0.01]*10).reshape(2, 5),
... bins=bins5, name='ds5')
>>> ds1 + ds5
Traceback (most recent call last):
[...]
ValueError: Datasets to add do not have same bin names
>>> f"bins ds1: {list(ds1.bins.keys())}, bins ds5: {list(ds5.bins.keys())}"
"bins ds1: ['e', 't'], bins ds5: ['E', 't']"
>>> bins6 = OrderedDict([('e', np.array([1, 2, 30])), ('t', np.arange(5))])
>>> ds6 = Dataset(np.arange(0, -10, -1).reshape(2, 5),
... np.array([0.01]*10).reshape(2, 5),
... bins=bins6, name='ds6')
>>> ds1 - ds6
Traceback (most recent call last):
[...]
ValueError: Datasets to subtract do not have the same bins
>>> same_coords(ds1, ds6)
False
>>> list(ds1.bins.keys()) == list(ds6.bins.keys())
True
>>> np.array_equal(ds1.bins['e'], ds6.bins['e'])
False
>>> np.array_equal(ds1.bins['t'], ds6.bins['t'])
True
Multiplication and division
Multiplication and division are possible between a Dataset and a
scalar (numpy.generic), a numpy.ndarray and another
Dataset.
The operator to use for multiplication is * and for division /.
Restrictions in multiplication and division with a numpy.ndarray are
handled by NumPy.
The multiplication or division of 2 Dataset can be done if
both values have the same shape (
consistent_datasets)bins conditions (
same_coords):EITHER the second
Datasetdoes not have any binsOR bins are the same, i.e. have the same keys and bins values
Division by zero, nan or inf are handled by NumPy and return a RuntimeWarning from NumPy (only in the zero case).
Example multiplication of a scalar value
>>> ds1m10 = ds1 * 10
>>> ds1m10.__class__
<class 'valjean.eponine.dataset.Dataset'>
>>> np.array_equal(ds1m10.value, [[0, 10, 20, 30, 40],
... [50, 60, 70, 80, 90]])
True
>>> np.array_equal(ds1m10.error, np.array([3.]*10).reshape(2, 5))
True
>>> same_coords(ds1m10, ds1)
True
As expected it acts on the value and on the error. Bins are unchanged.
You can also put the scalar value first:
>>> ds1m10_inv = 10 * ds1
>>> np.array_equal(ds1m10.value, ds1m10_inv.value)
True
>>> np.array_equal(ds1m10.error, ds1m10_inv.error)
True
>>> same_coords(ds1m10, ds1m10_inv)
True
Example of division of a numpy.ndarray
>>> ds1da = ds1 / a
>>> ds1da.name == ds1.name
True
>>> ds1da.what == ds1.what
True
>>> np.array_equal(ds1da.value, [[0., 0.01, 0.02, 0.03, 0.04],
... [0.05, 0.06, 0.07, 0.08, 0.09]])
True
>>> np.array_equal(ds1da.error, np.array([0.003]*10).reshape(2, 5))
True
>>> same_coords(ds1da, ds1)
True
a and ds1 have the same shape so everything is fine.
>>> ds1 / b
Traceback (most recent call last):
...
ValueError: operands could not be broadcast together with shapes (2,5) (10,)
If the shapes are not the same NumPy raises an exception.
If the numpy.ndarray: contains 0, nan or inf, NumPy deals
with them. It sends a RunningWarning about the division by zero.
>>> c = np.array([[2., 3., np.nan, 4., 0.], [1., np.inf, 5., 10., 0.]])
>>> ds1 / c
class: <class 'valjean.gavroche.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
name: ds1, with shape (2, 5),
value: [[0. 0.33333333 nan 0.75 inf]
[5. 0. 1.4 0.8 inf]],
error: [[0.15 0.03333333 nan 0.075 inf]
[0.3 0. 0.06 0.03 inf]],
bins: OrderedDict([('e', array([1, 2, 3])), ('t', array([0, 1, 2, 3, 4]))])
>>> # prints *** RuntimeWarning: divide by zero encountered in true_divide
Example of multiplication or division of another Dataset
>>> ds1m2 = ds1 * ds2
>>> ds1m2
class: <class 'valjean.gavroche.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape (2, 5),
value: [[ 0 21 44 69 96]
[125 156 189 224 261]],
error: [[6. 6.31268564 6.64830806 7.00357052 7.37563557]
[7.76208735 8.16088231 8.57029754 8.98888202 9.4154129 ]],
bins: OrderedDict([('e', array([1, 2, 3])), ('t', array([0, 1, 2, 3, 4]))])
>>> ds1m2.what
'spam*spam'
>>> same_coords(ds1m2, ds2)
True
>>> ds1o2 = ds1 / ds2
>>> ds1o2
class: <class 'valjean.gavroche.dataset.Dataset'>, data type: <class 'numpy.ndarray'>
shape (2, 5),
value: [[0. 0.04761905 0.09090909 0.13043478 0.16666667]
[0.2 0.23076923 0.25925926 0.28571429 0.31034483]],
error: [[0.015 0.01431448 0.01373617 0.01323926 0.01280492]
[0.01241934 0.01207231 0.01175624 0.01146541 0.0111955 ]],
bins: OrderedDict([('e', array([1, 2, 3])), ('t', array([0, 1, 2, 3, 4]))])
>>> ds1o2.what
'spam/spam'
In both cases the error is calulated considering both datasets are independent,
so quadratically 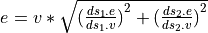 .
.
The same restictions on bins as for addition and subtraction are set for
multiplication and division, same AssertError are raised, see
Example of addition or subtraction of another Dataset.
About the division by 0, nan or inf, it acts like in the
multiplication or division by a numpy.ndarray, see
Example of division of a numpy.ndarray (warnings and nan, inf, etc.)
>>> np.isnan((ds1/ds1).value[0][0])
True
>>> np.isinf(((ds1+1)/ds1).value[0][0])
True
Comparisons
There are many possible criteria to compare Dataset objects. For
instance, one might consider that datasets are equal if they hold the same
values, regardless of the bins or the errors, or one might consider that even
metadata such as names should influence the comparison. For these reasons, the
use of the == operator (as defined in __eq__) is not recommended.
To accommodate all these possibilities, there is a method called
equals, which can be customized to include or exclude various
parts of the Dataset from the comparison. For example:
>>> ds = Dataset(value=np.arange(20, 30).reshape(2, 5),
... error=np.array([0.4]*10).reshape(2, 5),
... bins=bins, name='dataset', what='spam')
>>> ds2 = ds.copy()
>>> ds.equals(ds2)
True
By default, equals compares the following dataset attributes:
shape
value
error
bins
Any of these attributes can be excluded from the comparison using the flag of the same name. For instance:
>>> ds2.value = ds.value + 1
>>> ds.equals(ds2)
False
>>> ds.equals(ds2, value=False) # we exclude value
True
Likewise, name and what can be included in the comparison:
>>> ds2 = ds.copy()
>>> ds2.name = 'sblinda'
>>> ds.equals(ds2) # with default options, we don't compare ``name``
True
>>> ds.equals(ds2, name=True) # but we can!
False
NaN values are not treated as equal by default:
>>> ds = Dataset(value=np.arange(20, 30).reshape(2, 5),
... error=np.array([0.4]*10).reshape(2, 5),
... bins=bins, name='dataset', what='spam')
>>> ds.error[0, 2] = np.nan
>>> ds.equals(ds)
False
>>> ds.equals(ds, equal_nan=True)
True
Indexing and slicing
It is only possible to get a slice of a dataset, getting an Dataset at a given index is not possible (for dimensions consistency reasons). Requiring a given index can then be done using a slice.
Getting a subset of the Dataset
Time is the second dimension, to remove first and last bins the usual slice is
[1:-1], the first dimension, energy, is conserved, so its slice is [:].
The slice to apply is then [:, 1:-1].
>>> ds1sltfl = ds1[:, 1:-1]
>>> ds1sltfl.__class__
<class 'valjean.eponine.dataset.Dataset'>
>>> np.array_equal(ds1sltfl.value, [[1, 2, 3], [6, 7, 8]])
True
>>> np.array_equal(ds1sltfl.error, np.array([0.3]*6).reshape(2, 3))
True
>>> list(ds1sltfl.bins.keys()) == list(ds1.bins.keys())
True
>>> np.array_equal(ds1sltfl.bins['e'], ds1.bins['e'])
True
>>> np.array_equal(ds1sltfl.bins['t'], ds1.bins['t'])
False
>>> np.array_equal(ds1sltfl.bins['t'], [1, 2, 3])
True
>>> ds1sltfl.name == ds1.name
True
Slicing is also applied on bins.
Warning
Requiring a slice when there are not enough elements on the dimension give empty arrays.
For example: removing first and last bin in energy on ds1. The slice is
[1:-1, :] in that case, but ds1 has only 2 bins in energy.
>>> ds1slefl = ds1[1:-1, :]
>>> ds1slefl.value.shape == (0, 5)
True
>>> np.array_equal(ds1slefl.value, np.array([]).reshape(0, 5))
True
>>> np.array_equal(ds1slefl.error, np.array([]).reshape(0, 5))
True
>>> np.array_equal(ds1slefl.bins['t'], ds1.bins['t'])
True
>>> np.array_equal(ds1slefl.bins['e'], ds1.bins['e'])
False
>>> np.array_equal(ds1slefl.bins['e'], [2])
True
Note that in this case, as bins are in reality the edges of the bins, so we have N+1 values in the bins compared to the value/error where we have N values. Slicing then give one value in energy bins, so unusable here (it would be empty if we have values and centers of bins instead of edges of bins).
All dimensions have to be present in the slice
Use : for the untouched dimensions. Number of , has to be dimension -1.
Let’s consider a new Dataset, with 4 dimensions:
>>> bins2 = OrderedDict([('e', np.arange(4)), ('t', np.arange(3)),
... ('mu', np.arange(3)), ('phi', np.arange(5))])
>>> ds6 = Dataset(np.arange(48).reshape(3, 2, 2, 4),
... np.array([0.5]*48).reshape(3, 2, 2, 4),
... bins=bins2, name='ds6')
>>> ds6.value.ndim == 4
True
To remove first bin on energy dimension and last bin on phi dimension, the
slice to be used is: [1:, :, :, :-1].
>>> ds6_1 = ds6[1:, :, :, :-1]
>>> ds6.value.shape == (3, 2, 2, 4)
True
>>> ds6_1.value.shape == (2, 2, 2, 3)
True
>>> np.array_equal(ds6_1.value, [[[[16, 17, 18], [20, 21, 22]],
... [[24, 25, 26], [28, 29, 30]]],
... [[[32, 33, 34], [36, 37, 38]],
... [[40, 41, 42], [44, 45, 46]]]])
True
>>> np.array_equal(ds6_1.error, np.array([0.5]*24).reshape(2, 2, 2, 3))
True
>>> list(ds6_1.bins.keys()) == list(ds6.bins.keys())
True
>>> np.array_equal(ds6_1.bins['e'], ds6.bins['e'][1:])
True
>>> np.array_equal(ds6_1.bins['t'], ds6.bins['t'])
True
>>> np.array_equal(ds6_1.bins['mu'], ds6.bins['mu'])
True
>>> np.array_equal(ds6_1.bins['phi'], ds6.bins['phi'][:-1])
True
If we only want the second bin in time keeping all bins in energy and direction
angles, the slice is [:, 1:2, :, :].
>>> ds6_2 = ds6[:, 1:2, :, :]
>>> ds6_2.value.shape == (3, 1, 2, 4)
True
>>> np.array_equal(ds6_2.value, [[[[8, 9, 10, 11], [12, 13, 14, 15]]],
... [[[24, 25, 26, 27], [28, 29, 30, 31]]],
... [[[40, 41, 42, 43], [44, 45, 46, 47]]]])
True
>>> list(ds6_2.bins.keys()) == list(ds6.bins.keys())
True
>>> all(x.size == y+1
... for x, y in zip(ds6_2.bins.values(), ds6_2.value.shape)) == True
True
>>> np.array_equal(ds6_2.bins['t'], [1, 2])
True
Bins are changed accordingly.
Warning
Comparison to NumPy: index and ellipsis are other slicing possibilities
on numpy.ndarray (see numpy indexing for current version of
NumPy), but they are disabled here to avoid confusions. Errors are raised
if required.
>>> ds1[1]
Traceback (most recent call last):
[...]
TypeError: Index can only be a slice or a tuple of slices
>>> ds6_2e = ds6[:, 1, :, :]
Traceback (most recent call last):
[...]
TypeError: Index can only be a slice or a tuple of slices
>>> ds6e = ds6[1:, ..., :-1]
Traceback (most recent call last):
[...]
TypeError: Index can only be a slice or a tuple of slices
It also need to have the same dimension as the value:
>>> ds6_2e = ds6[:, 1:2]
Traceback (most recent call last):
[...]
ValueError: len(index) should have the same dimension as the value numpy.ndarray, i.e. (# ',' = dim-1).
':' can be used for a slice (dimension) not affected by the selection.
Slicing is only possible if ndim == 1
A single slice is only possible for arrays for dimension 1:
>>> ds6_2f = ds6[1:2]
Traceback (most recent call last):
[...]
ValueError: len(index) should have the same dimension as the value numpy.ndarray, i.e. (# ',' = dim-1).
':' can be used for a slice (dimension) not affected by the selection.
Slicing is only possible if ndim == 1
>>> ds7 = Dataset(value=np.arange(48), error=np.array([1]*48))
>>> ds7.ndim
1
>>> ds7.shape
(48,)
>>> ds7_extract = ds7[20:24]
>>> np.array_equal(ds7_extract.value, [20, 21, 22, 23])
True
Warning
Slicing can also only be applied on numpy.ndarray, not on
numpy.generic:
>>> ds8 = Dataset(value=100, error=1)
>>> ds8[0:1]
Traceback (most recent call last):
[...]
TypeError: [] (__getitem__) can only be applied on numpy.ndarrays
Masked datasets
In some case it can be useful to mask some elements of a dataset. This functionality is provided by the numpy masked arrays module. In the case of datasets, once the mask is given it is applied to both the value and the error.
>>> mask = np.ma.masked_greater(ds1.value, 6).mask
>>> np.array_equal(mask, [[False, False, False, False, False],
... [False, False, True, True, True]])
True
>>> mds = ds1.mask(mask)
>>> np.ma.is_masked(ds1.value)
False
>>> np.ma.is_masked(mds.value)
True
>>> np.ma.is_masked(mds.error)
True
Bins and shape are kept.
>>> mds.shape == ds1.shape
True
>>> np.array_equal(mds.bins['e'], ds1.bins['e'])
True
>>> np.array_equal(mds.bins['t'], ds1.bins['t'])
True
The mask is propagated when performing operations on dataset.
>>> np.sum(ds1.value) == 45
True
>>> np.sum(mds.value) == 21
True
>>> sds = ds1 + mds
>>> np.ma.is_masked(sds.value)
True
>>> np.ma.is_masked(sds.error)
True
>>> np.sum(sds.value) == 42
True
- class valjean.eponine.dataset.Dataset(value, error, *, bins=None, name='', what='')[source]
Common class for data from all codes.
For the moment, units are not treated (removed).
Todo
Think about units. Possibility: using a units package from scipy.
Todo
How to deal with bins of N values (= center of bins)
- __init__(value, error, *, bins=None, name='', what='')[source]
Dataset class initialization.
- Parameters:
value (int or float or numpy.ndarray or numpy.generic) – array of N dimensions representing the values
error (int or float or numpy.ndarray or numpy.generic) – array of N dimensions representing the absolute errors
bins (collections.OrderedDict (str, numpy.ndarray)) – bins corresponding to value (named optional parameter)
name (str) – name of the dataset (used in test representation)
what (str) – name of the quantity represented by the dataset
- squeeze()[source]
Squeeze dataset: remove useless dimensions.
Squeeze is based on the shape and on the bins dim ??? To confirm… First squeeze bins, then arrays. Edges, if only one bin are not kept. Example: spectrum with one bin in energy (quite common)
- property shape
Return the data shape, as a read-only property.
- property ndim
Return the data dimension, as a read-only property.
- property size
Return the data size (total number of elements in the array), as a read-only property.
- data()[source]
Generator yielding objects supporting the buffer protocol that (as a whole) represent a serialized version of self.
>>> from valjean.fingerprint import fingerprint >>> vals = np.arange(10).reshape(2, 5) >>> errs = np.array([1]*10).reshape(2, 5) >>> ds = Dataset(vals, errs) >>> fingerprint(ds) 'a8411470d7766c543e90f0f38241dc918b9448d1b9d19b0a9b8b6c91f61944d0' >>> bins = OrderedDict([('bacon', np.arange(3)), ... ('egg', np.arange(5))]) >>> ds = Dataset(vals, errs, bins=bins) >>> fingerprint(ds) 'dfec4e6ac118d30b7a83cfc500fefaec94f93a76a024d7550732cd08fbfb0fee' >>> ds = Dataset(vals, errs, bins=bins, name='name') >>> fingerprint(ds) '441623c096bf917414013ebbfe345c66124f9a8dfbbd7051cf733ea20d7b651a' >>> ds = Dataset(vals, errs, bins=bins, name='name', what='what!') >>> fingerprint(ds) '9a42c91381d696c1af651665f84efc43efa14d9c471c4ac851239ed0779a48a0'
- mask(mask)[source]
Apply a mask to the Dataset value and error.
Value and error become in that case masked_arrays instead of usual arrays, calcuations are preserved, not using the masked elements.
- Return type:
- equals(other, *, shape=True, value=True, error=True, bins=True, name=False, what=False, equal_nan=False)[source]
Compare two datasets
- Parameters:
shape (bool) – whether dataset shapes should be compared
value (bool) – whether dataset values should be compared
error (bool) – whether dataset errors should be compared
bins (bool) – whether dataset bins should be compared
name (bool) – whether dataset names should be compared
what (bool) – whether dataset whats should be compared
equal_nan (bool) – whether NaN values should be considered equal; it works like the parameter with the same name in
numpy.array_equal
- Return type:
- Returns:
whether the two arrays are equal